Project
Quantitative understanding of a dynamic protein interaction network
Biological functions of a protein depend largely on it’s cellular localization and concentration. These parameters are even more crucial for regulatory proteins that drive hormone signaling pathways such as auxin. To understand the role of dynamic protein parameters in controlling auxin sensitivity and response, we are studying the auxin response proteins at their endogenous concentrations in Marchantia polymorpha. Using in-vivo confocal time-course imaging of knock-in lines and biochemical analysis of recombinant proteins we aim to uncover the quantitative aspects of the auxin response network.
Background
The plant hormone auxin steers various complex growth and developmental processes through an apparently simple nuclear signaling pathway that mostly acts via transcriptional gene regulation. The nuclear auxin signaling pathway (NAP) has only three set of components: the nuclear auxin receptor TRANSPORT INHIBITOR RESPONSE 1 (TIR1), the repressor Auxin/INDOLE-3-ACETIC ACID (Aux/IAA) and the AUXIN RESPONSE FACTOR (ARF) transcription factors. The liverwort model plant Marchantia polymorpha has a minimal nuclear auxin response pathway with only 1 TIR1, 1 Aux/IAA and 3 ARFs representing 3 different ARF classes present in Angiosperms. Taking advantage of this simple NAP, we are investigating the auxin response proteins at native concentrations in homologous knock-in translational fusion lines.
Aim of the project
ARFs are transcription factors that regulate gene transcription in response to auxin. Although ARFs have been functionally characterized in multiple species, information on native ARF expression dynamics during growth is scant. Using time course imaging on developing Marchantia polymorpha we are investigating endogenous ARF dynamics. Using molecular genetic tools and pharmacological treatments we are altering the native protein stoichiometry to understand the biological significance of changing ARF levels in various tissues and how this influences plant growth. Apart from this we are also looking for novel ARF interactors that can form complexes and influence ARF stability and functionality.
Results
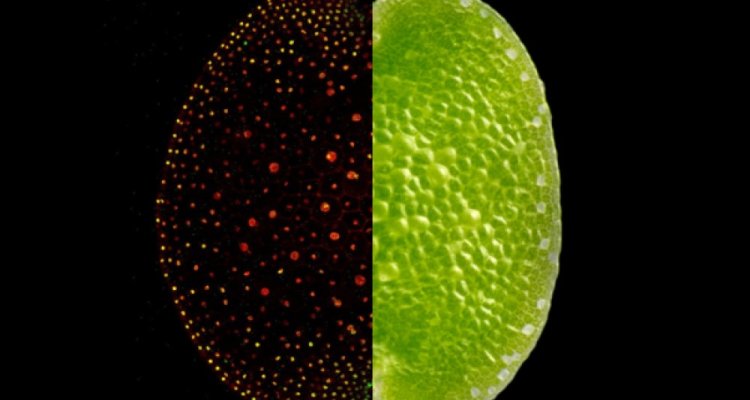
We have successfully mapped the native expression patterns of all auxin response proteins in Marchantia polymorpha. Currently, we are investigating further into the native protein complex stoichiometry, protein degradation rates etc. that might regulate the auxin response network.
Contact
Do you have a question about the quantitative and dynamic protein interaction networks, or would you like to join us as a student researcher? Please contact us.