Project
Microbial interactions in marine anoxic sediments: competition or teamwork?
Biodegradation of organic matter (OM) in marine sediments occurs through a sequence of reactions by a diverse population of aerobic and anaerobic microbes. Fermentative bacteria convert oligomers to a variety of products, mainly short chain fatty acids, CO2 and hydrogen. Further conversion through the action of secondary fermenters, sulfate-reducers, acetogens and methanogens produces the end products CO2, CH4 and H2S. When sulfate is absent, OM is mineralized exclusively to CO2 and methane.
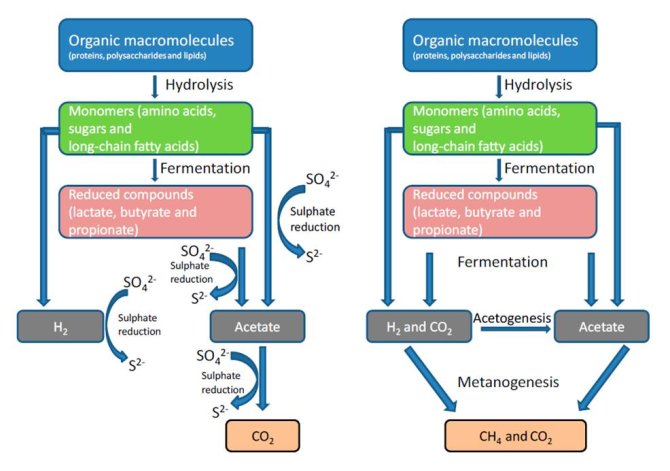
Recently it was demonstrated that in marine sediments hydrogen was produced during biodegradation and influenced the metabolism of both methanogens and sulfate-reducers. This has implications for overall carbondegradation pathways.
Our aim with this project is to better understand the microbial interactions and metabolic flexibility in coastalmarine sediments. In this manner, we propose to study microbial diversity and microbial interactions using bothcultivation-dependent and cultivation-independent techniques, and mixed-cultivation experiments mimicking insitu conditions.
We will identify, isolate and cultivate a variety of anaerobic microorganisms that are present in the sulfate, methane and sulfate/methane transition zones of coastal marine sediments. We aimed to demonstrate that syntrophic growth is a common mechanism for anaerobic degradation of short-chain fatty acids and C-1 compounds in OM rich marine sediments and thus is a significant process. Since it is still unclear which type of metabolism sulfate reducers encompass in the methane zone, and vice versa, we will focus on unraveling unusual interactions that occur in marine sediments. Presence and in-situ activity of microbial consortia will be demonstrated with culture-independent 13C-Stable Isotope Probing (SIP). Depending on the metabolic pathways employed, the label ends up in different metabolic products and in different carbon positions. Thus, quantification of 13C-labeled metabolic products allows additional characterization of microbial functionality, because specific metabolic pathways can be directly linked to distinct microbial populations when it is combined with nucleic acid based SIP.
The mechanisms and microorganisms studied will considerably increase our understanding of biogeochemical zonation due to terminal electron acceptor availability and competition for substrates.
Keywords: Sulfate-reducing bacteria (SRB), methanogens, anoxic marine sediments, syntrophy, DGGE, FISH, RT-qPCR, 13C-stable isotope probing (SIP).
If you have further questions, or want to do a thesis / internship on this subject, please contact me.