
Software
Pedimap
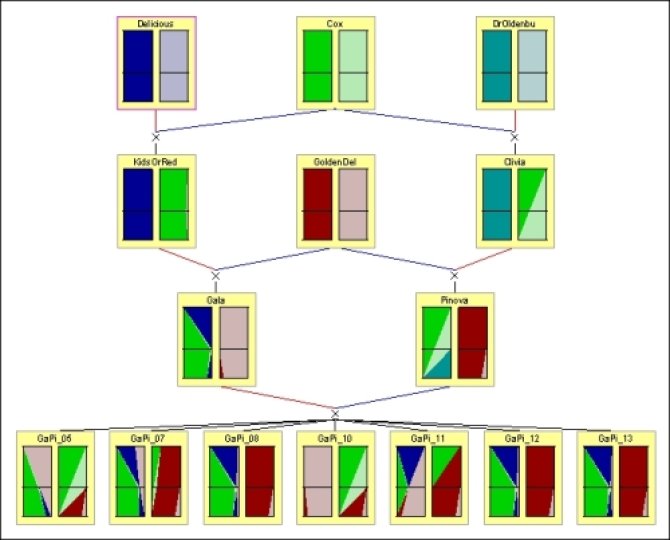
Pedimap is a tool for exploring and visualizing the flow of phenotypes and alleles (observed or based on Identity-by-Descent calculations) through pedigrees.
Pedimap is a useful stand-alone tool to check marker data and to understand the relation between phenotype and genotype. Further, Pedimap can be used to display the results calculated by the FlexQTL software developed by Dr Marco Bink of WUR – Biometris. Pedimap can quickly highlight problematic marker scores and display the calculated Identity-by-Descent information. Pedimap and FlexQTL are designed to work together.
Funding
Pedimap was developed at WUR – Plant Breeding. Its development was started in the framework of the European HiDRAS (www.hidras.unimi.it) project and continued within USDA-NIFA-RosBREED (www.rosbreed.org) and EU-FruitBreedomics (www.fruitbreedomics.com) projects.