
Nieuws
Pascal Duenk wins Overall best poster prize at ICQG6
During the 6th International Conference on Quantitative Genetics (ICQG6), Pascal Duenk from Wageningen University & Research won the overall best poster prize. In total, almost 200 posters were presented at ICQG6, which was held online over the course of two weeks, between November 2 and 12.
In his poster presentation, Duenk presented the last chapter of his PhD thesis, which he defended on 13 March 2020.
Purebred-crossbred genetic correlation
The purebred-crossbred genetic correlation (rpc) is a key parameter in breeding programs that rely on crossbreeding, such as pig and poultry breeding programs. This correlation determines to what extend genetic progress in purebred breeding populations is transferred to their crossbred descendants in commercial farms.
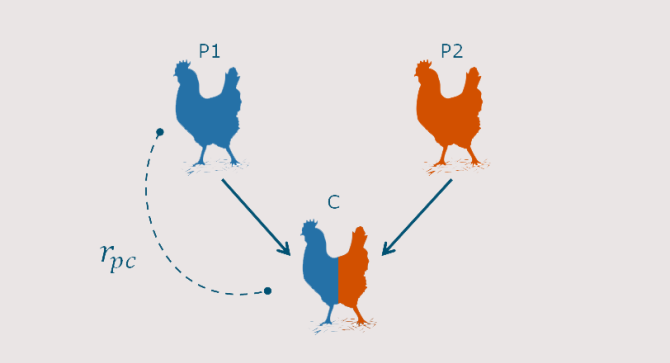
Estimating the rpc requires measuring performance of crossbred animals, which is costly, and not necessarily done routinely within all breeding programs. Pascal developed formulas that accurately predict an upper and lower limit of the purebred-crossbred correlation, using only information obtained from purebred animals.
Non-additive effects
The rpc can be lower than 1.0 due to genotype by environment interactions and non-additive effects. Duenk’s formulas predict the contribution of non-additive effects to the rpc, based on the genetic correlation between the parental breeds of the crossbred. Using simulations, he showed that assuming a dominance model yields a lower limit, while an epistatic model yields an upper limit for the purebred-crossbred correlation.
This work is currently under review at Genetics Selection Evolution.