Project
Exploring epistatic interactions in natural genetic variation for photosynthesis
Introduction
By 2050, it is expected that the worldwide agricultural crop productivity will need to increase by 60% to meet future demands for a growing population, but annual yield increases are stagnating despite high-tech plant breeding approaches, Photosynthesis efficiency in plants is an underexploited trait to improve our crop species, but recently published papers report biomass increases of 15%-40% in plants that had their photosynthesis efficiency altered when grown under conditions normal to crop plants. Improving photosynthesis may therefore become a key trait in the coming decades. However, it is generally not used by plant breeders, due to the complexity of phenotyping genetic mapping populations. Furthermore, its complex genetic architecture constrains pinpointing the causal genes or genetic interactions that may contribute to this trait. At the Laboratory of Genetics we have developed genetic tools, analyses and populations in order to overcome such difficulties.
Aim
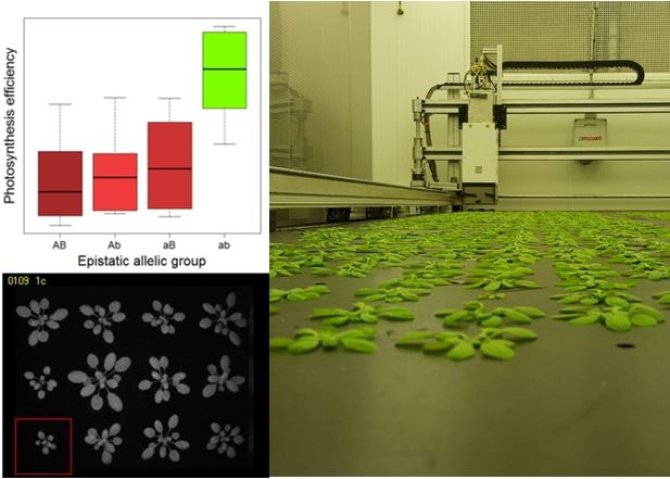
The aim of this research is to use novel genetic resources, in the form of chromosome substitution libraries (CSLs), and new, genome-wide, epistatic analysis software to reveal functional genetic variation that is otherwise undetectable in conventional mapping approaches. Of special interest is epistasis, interaction between genetic loci (Fig. 1), which is expected to play a big role in photosynthesis due the total number of genes that are involved but remains elusive. The ultimate goal of this project is to discover new genes and traits that may contribute towards improving photosynthesis efficiency, and eventually the required increase in crop productivity.
Approach
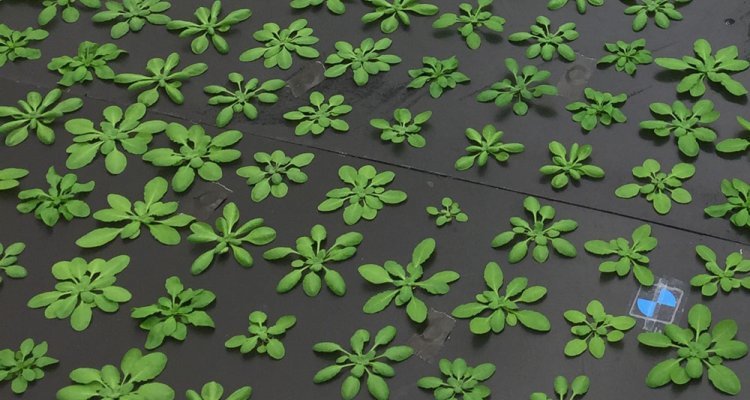
The methods that are employed in this study are firstly genetic mapping, in CSLs or using novel algorithm that allow the detection of epistasis in large GWAS populations. Phenotyping is performed in a state of the art high-throughput phenotyping platform, which can assess daily photosynthesis of up to 1440 plants, five times a day, and make use of different photosynthetic conditions (Fig. 2). Thereafter, fine mapping should lead to candidate gene discovery which will then be the subjected to expression (q/RT PCR), allelic and transgenic (e.g. by using CRISPR-cas9, T-DNA mutant lines) analysis. If possible, biomass and/or biochemical analyses will be performed to directly determine the impact of photosynthesis on plant productivity and physiology (Fig. 3).
A typical thesis project will contain both a genetic analytical and a molecular genetics component, although the size of each of these may vary according to the state of the research and the student’s interests.
Student Opportunities
- New thesis projects are available from Autumn 2019 and onwards
- Are you interested? Contact Mark.Aarts@wur.nl