Project
Exploring metabolic capabilities of novel anaerobes
Introduction
Novel anaerobic bacteria are everywhere! From the domestic to the most extreme environments, from waste waters to the human gut, novel bacteria are constantly discovered. Such microorganisms although they get isolated, they still remain fully unknown. A great potential that just waits to be explored. So we must be ready to explore new capabilities, new ideas and new perspectives!
With innovative technologies and recent methods, we can now dive in the genomes and the proteomes of the microorganisms achieving a detailed understanding of each novel bacteria. We can draw its metabolic profile and compare it with other species to identify and understand new properties.
This project includes the steps of a full research strategy with a combination of physiological characterization with laboratory experiments and genomic, proteomic and metabolic reconstruction with in silico analysis. The incredible potential of the bioinformatics is verified in the lab.
Research
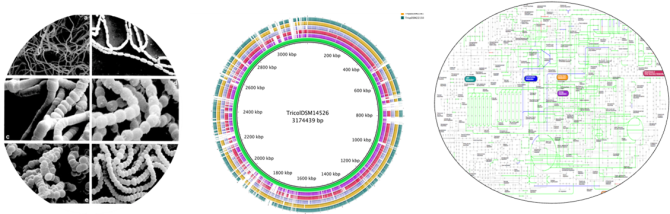
Currently, we are working with a bacteria isolated from NASA in the Chilian Patagonia from penguin gut. It grows in -5oC creating a blanket of mucous substance to withstand cold. Another case of bacteria of interest is one that produces 1-3 propanediol, which is a major compound for plastic biosynthesis. There is even a complete novel genus (Trichococcus spp.) that must be completely analysed and determined its purpose in the bacteria kingdom. The species of this novel genus grow in a great range of temperature from 0-45 degrees and can degrade a broad spectrum of carbohydrate substrates.
Trichococcus species are so identical, but so different at the same time. They all share around 100% 16 rRNA gene similarity. This is a highly rare case for microorganisms. On the other hand, their metabolic profile is highly divergent. These bacteria as they grow in one day can contribute in new compounds and applications for biotechnology and answer evolutionary questions regarding their own evolution.
Project
This project includes a union of wet and dry experiments. The novel bacteria can be either analyzed already for their interesting physiology or also a “genome-guided” physiological analysis. For describing the bacteria, genome analysis is required combined with differential proteomics and metabolic profiling. The combination of omics studies will lead in pathway reconstruction and optimization of biotechnological processes.
This project includes:
· Laboratory experiments based on the physiology of microorganisms
· Laboratory proteome extraction methods
· Laboratory RNA extraction methods
· Comparative genomics and genome analysis
· Comparative transcriptomics and transcriptome analysis
· Comparative proteomic and protein analysis
· Metabolic pathways analysis
· Psychrophilic enzymes and substance analysis