
Project
Protein balance of Lactococcus lactis in cheese production
Background
In dairy fermentations lactic acid bacteria are used as starter cultures. Especially starter cultures employed in cheese manufacturing, encounter many different environmental conditions during their production and during their application in cheese manufacturing. To optimally perform, microorganisms have to continuously adapt to these changing conditions. Bacteria adjust their physiology to maximize their fitness to the condition they are in, which often comes at the cost of being less fit under different conditions. The fitness of a microorganism under different conditions is determined by their rate of adaptation, which depends on the degradation, production and renewal of their proteome. However, this adaptation and turnover of the proteome is constrained by the availability of resources and/or cellular energy levels, which results in trade-offs during optimized growth followed by adaptation (the optimization of one trait can only occur at the cost of another).
Objectives
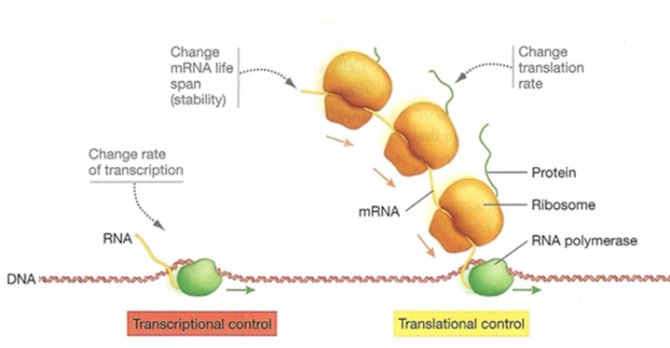
In this project, we want to understand the regulation of the protein synthesis and turnover in Lactococcus lactis under several different conditions using whole genome proteomics. In addition, we focus on enzymes involved in cheese flavour formation to investigate how trade-offs can influence the performance of cheese starter cultures.
Methodology
Bacterial culturing, RNA isolation, protein isolation, transcriptomics, proteomics, bioinformatics
Requirements
Students pursuing a degree in Molecular Life Sciences, Biotechnology, Biology, Bioinformatics or related discipline.
Background in either molecular biology/biochemistry (previous experience with microbiological culturing is preferred) or in bioinformatics.
For this project, an interest in proteins/proteomics is important. However, if you like the project but you are more interested in evolutionary biology, data-modelling or enzyme catalysis, you can also contact us.
Contact information
Berdien van Olst Berdien.vanolst@wur.nl