Project
Assessment of the role of horizontal gene tranfer in the bacteril adaptation to halogenated compounds in the environment
The traditional view on evolution has been that small mutations may occur in a certain organism and if beneficial, give the organism an increased fitness leading to this organism having a higher number of offspring; having the new beneficial mutations.
More and more small mutations may accumulate in different lineages leading them to gradually differentiate into new species. A popular way to depict this has been in the form of a tree were different organisms are grouped after relatedness, mainly based on morphology (fig 1.)
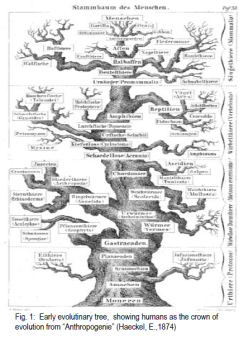
Modern DNA based tools however have enabled scientists to look into the evolutionary processes at the gene level making it clear that evolution is a far more dynamic and complex process than previously thought. It is now clear that genes are not only transferred laterally from parent to offspring but also horizontally between organisms of the same or different species or even from one kingdom to another (1, 2, 4, 8, 10, 11). This has lead some investigators to suggest that the traditional tree of life should be replaced by a far more complex web of life (fig 2 (5, 6)).
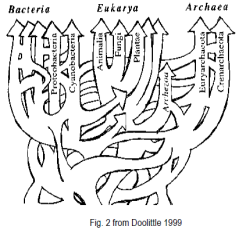
Well known examples of this are the appearance of multi-resistant pathogenic bacteria, which is of great concern to the medical industry.
Early research on horizontal gene transfer, focused on the transfer of plasmids, which are extra chromosomal pieces of DNA capable of independent replication. Today it is a well established fact that plasmids are transferred between different bacteria and in this way, horizontally spreading the genes they carry, for example resistance to certain antibiotics. As more and more bacteria are being sequenced, the evidence has increased that horizontal gene transfer plays a crucial role in the evolution of bacterial genomes, due to transfer of genetic elements that integrates into the genome (3, 14, 16, 18)
Aim
The aim of this project is to explore the role of horizontal gene transfer in microbial adaptation to xenobiotics, using reductive dehalogenation (RD) as a model system.
Research
Reductive dehalogenation
Halogenated hydrocarbons are widely used in the industry and private homes as for example biocides, solvents, degreasers, flame retardants and in polymer production, resulting in the release of large quantities into the environment. Due to the toxicity of many of these compounds, are they regarded as a great environmental concern (19, 21). It has been demonstrated that several bacteria are able to use these compounds as a growth substrate, both under aerobic and anaerobic conditions. Especially the latter has attracted attention, since it appears that polyhalogenated compounds are more readily dehalogenated under anaerobic than aerobic conditions (7, 19).
Anaerobic reductive dehalogenation is believed to be of ancient evolutionary origin, derived from the degradation of naturally produced haloorganics. This view is supported by the presence of highly conserved motifs in diverse and widely distributed reductive dehalogenases (RD) (19).
There is growing evidence that dehalogenases are subjected to horizontal gene transfer, based on the observation that transposases and other genes known to be involved in mobilization of DNA often are found close to dehalogenase coding genes (12, 20).
Recently the full genomes of three strains known to be capable of anaerobic dehhalogenation, i.e. Dehalococcoides ethenogens strain 195; Dehalococcoides strain CBDB1 and Desulfitobacterium hafnience strain Y51 have been published.
They were shown to contain between 2 and 32 putative RDs, and in all three strains at least one RD was located on putative mobile genetic elements (MGE). Putative MGEs were identified by either; presence of genes known to be associated with MGEs; anomalities in GC content; frequency of dinucleotides (also called Karlin signature, (13, 14) ); atypical trinucleotides; tetra nucleotides or a combination of these. (14, 16-18)
Several genomes of dehalogenating bacteria are currently being sequenced, and draft sequences of some of these are available for download for example Dehalococcoides sp. BAV1 and Desulfitobacterium hafnience DCB2 from the Joint Genome Institute.
Two recent studies on putative transposons in Desulfitobacterium sp. strain Y51 and Desulfitobacterium hafnience strain TCE1, revealed the presence of the transposons existing in both an integrated and a circularized intermediate free form in both strains. The transposons in both strains did consist of pceA containing gene-clusters flanked by two insertion sequences (9, 15). The formation of a circularized intermediate following excision of the transposon is believed to be a general step in the movement of transposons (4).
The ability of Desulfitobacterium sp. strain Y51 to degrade tetrachlroroethene (PCE), were lost in 15 out of 18 investigated colonies after 56 generations growth in a non selective medium. The loss of the ability to degrade PCE, were shown to be accompanied by a deletion of either almost the entire MGE or, parts of this (9). These findings indicate that, at least some MGEs associated with halorespiration, may be very mobile. The rapid loss of the ability to degrade PCE, under non selective conditions, may suggest a pattern were the MGE are lost from the majority of the population under nonselective conditions and recruited when PCE are present in the environment. This hypothesis is supported by the report of several cases of increased conjugation frequency of transposons coding for resistance to tetracycline, when tetracycline were present in the medium (4).
Current work is focused on identifying mobile elements in bacteria known to be involved in degradation of chlorinated hydrocarbons, and developing tools to follow these processes in simple model systems.
References
1. Bertram, J., M. Stratz, and P. Durre. 1991. Natural Transfer of Conjugative Transposon Tn916 between Gram-Positive and Gram-Negative Bacteria. Journal of Bacteriology 173:443-448.
2. Buchananwollaston, V., J. E. Passiatore, and F. Cannon. 1987. The Mob and Orit Mobilization Functions of a Bacterial Plasmid Promote Its Transfer to Plants. Nature 328:172-175.
3. Burrus, V., and M. K. Waldor. 2004. Shaping bacterial genomes with integrative and conjugative elements. Research In Microbiology 155:376-386.
4. Churchward, G. 2002. Conjugative Transposons and Related Mobile Elements. American Society for Microbiology, Washington DC.
5. Doolittle, W. F. 1999. Phylogenetic classification and the universal tree. Science 284:2124-2128.
6. Doolittle, W. F., Y. Boucher, C. L. Nesbo, C. J. Douady, J. O. Andersson, and A. J. Roger. 2003. How big is the iceberg of which organellar genes in nuclear genomes are but the tip? Philosophical Transactions of the Royal Society of London Series B-Biological Sciences 358:39-57.
7. El Fantroussi, S., H. Naveau, and S. N. Agathos. 1998. Anaerobic dechlorinating bacteria. Biotechnology Progress 14:167-188.
8. Frost, L. S., R. Leplae, A. O. Summers, and A. Toussaint. 2005. Mobile genetic elements: The agents of open source evolution. Nature Reviews Microbiology 3:722-732.
9. Futagami, T., Y. Tsuboi, A. Suyama, M. Goto, and K. Furukawa. 2006. Emergence of two types of nondechlorinating variants in the tetrachloroethene-halorespiring Desulfitobacterium sp strain Y51. Applied Microbiology And Biotechnology 70:720-728.
10. Grohmann, E., G. Muth, and M. Espinosa. 2003. Conjugative plasmid transfer in gram-positive bacteria. Microbiology and Molecular Biology Reviews 67:277-+.
11. Heinemann, J. A., and G. F. Sprague. 1989. Bacterial Conjugative Plasmids Mobilize DNA Transfer between Bacteria and Yeast. Nature 340:205-209.
12. Janssen, D. B., I. J. T. Dinkla, G. J. Poelarends, and P. Terpstra. 2005. Bacterial degradation of xenobiotic compounds: evolution and distribution of novel enzyme activities. Environmental Microbiology 7:1868-1882.
13. Karlin, S. 1998. Global dinucleotide signatures and analysis of genomic heterogeneity. Current Opinion In Microbiology 1:598-610.
14. Kube, M., A. Beck, S. H. Zinder, H. Kuhl, R. Reinhardt, and L. Adrian. 2005. Genome sequence of the chlorinated compound respiring bacterium Dehalococcoides species strain CBDB1. Nature Biotechnology 23:1269-1273.
15. Maillard, J., C. Regeard, and C. Holliger. 2005. Isolation and characterization of Tn-Dha1, a transposon containing the tetrachloroethene reductive dehalogenase of Desulfitobacterium hafniense strain TCE1. Environmental Microbiology 7:107-117.
16. Nonaka, H., G. Keresztes, Y. Shinoda, Y. Ikenaga, M. Abe, K. Naito, K. Inatomi, K. Furukawa, M. Inui, and H. Yukawa. 2006. Complete genome sequence of the dehalorespiring bacterium Desulfitobacterium hafniense Y51 and comparison with Dehalococcoides ethenogenes 195. Journal Of Bacteriology 188:2262-2274.
17. Regeard, C., J. Maillard, C. Dufraigne, P. Deschavanne, and C. Holliger. 2005. Indications for acquisition of reductive dehalogenase genes through horizontal gene transfer by Dehalococcoides ethenogenes strain 195. Applied And Environmental Microbiology 71:2955-2961.
18. Seshadri, R., L. Adrian, D. E. Fouts, J. A. Eisen, A. M. Phillippy, B. A. Methe, N. L. Ward, W. C. Nelson, R. T. Deboy, H. M. Khouri, J. F. Kolonay, R. J. Dodson, S. C. Daugherty, L. M. Brinkac, S. A. Sullivan, R. Madupu, K. T. Nelson, K. H. Kang, M. Impraim, K. Tran, J. M. Robinson, H. A. Forberger, C. M. Fraser, S. H. Zinder, and J. F. Heidelberg. 2005. Genome sequence of the PCE-dechlorinating bacterium Dehalococcoides ethenogenes. Science 307:105-108.
19. Smidt, H., and W. M. de Vos. 2004. Anaerobic microbial dehalogenation. Annual Review Of Microbiology 58:43-73.
20. Springael, D., and E. M. Top. 2004. Horizontal gene transfer and microbial adaptation to xenobiotics: new types of mobile genetic elements and lessons from ecological studies. Trends In Microbiology 12:53-58.
21. van Pee, K.-H., and S. Unversucht. 2003. Biological dehalogenation and halogenation reactions. Chemosphere 52:299.
Student projects available
There are several exciting possibilities for doing a master project in the field of bacterial evolution and degradation of xenobiotics.
Hereunder are shortly mentioned a few examples of possible projects, if you are interested in doing one of these, have other ideas or questions then do not hesitate to contact me by mail: Thomas Kruse
Transposons guide to hitchhiking in the microbial world
Transposons are small genetic elements, with the ability to integrate themselves in the genome and later excise them self and jump to another place in the genome.
Transposons can vary in size from very small transposons that only contain the information necessary for integrating in the genome to large conjugative transposons