Nodulation Engineering (Geurts Group)
The above title is a citation of a Science paper in 2016 (Stockstad, 2016, Science 353:12225-7) that reflected the massive expectations researchers have about engineering nitrogen-fixing root nodules on crops. Such root nodules are well known from legumes and are the subject of study for over a hundred years.
However, not only legumes possess the trait. Outside the legume family, root nodules occur in 9 taxonomic lineages. Among these is Parasponia, a small genus in the Cannabis family, representing five tropical tree species. Parasponia trees can form nitrogen-fixing root nodules with the same rhizobium bacteria that also can nodulate legume plants.
Genomics and molecular genetic studies demonstrated that the nitrogen-fixing nodulation trait in legumes and Parasponia have a shared evolutionary origin. We use Parasponia as a comparative system to legumes to identify the critical genetic adaptations allowing plants to form nitrogen-fixing root nodules. Subsequently, this information is used to engineer crop plants, of which cassava is our main target.
The Nitrogen Fix – Few projects in plant biotechnology are harder, or promise a greater payoff than enabling crops to make their own nitrogen fertilizer.
Topics:
Comparative bioinformatics to identify genes that correlate with nodulation trait
By using a phylogenomic approach we discovered three critical genes that associate with nodulation. However, these genes may reflect only a tip of the iceberg. Ongoing genome sequencing initiatives allow high resolution comparisons of genomes and expression data of related nodulating and non-nodulating plant species. Besides gene discovery associated with symbiosis, we also aim to identify cis-regulatory elements regulating symbiotic genes that differ between nodulating and non-nodulating plants. In this way adaptations in nodulation genes and promotor regions can be discovered.
In a research thesis you will come familiar with bioinformatic tools and methods such as comparative genomics, working on the Linux command line, and programming in R or Python.
Design-Built-Test of engineering constructs
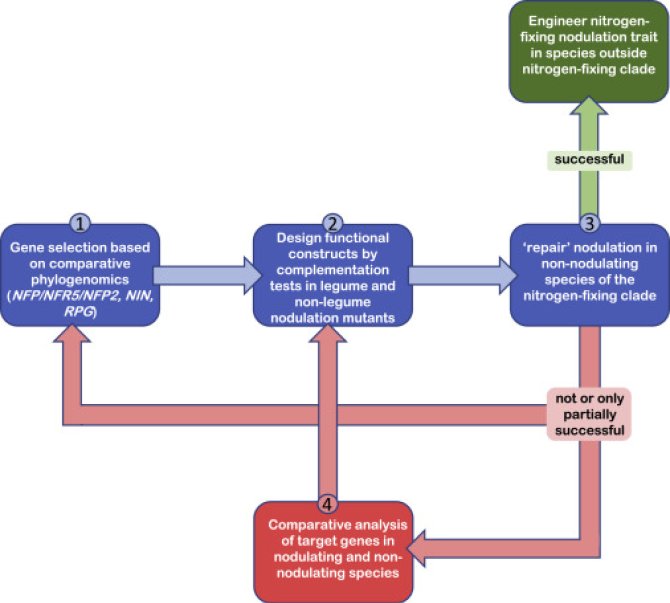
To successfully engineer a complex genetic trait in crops, it is essential to have a fast and efficient cycle of designing engineering constructs, built these constructs, and test them in vivo. The design of novel gene constructs is guided by bioinformatic analysis of nodulating and non-nodulating plant species. An important tool in our engineering strategy also involves the design of efficient marker gene constructs for discrete steps in rhizobium-induced nodule formation. These include markers specific for rhizobium-induced signalling, developmental markers to discriminate between nodule and lateral root organogenesis, and markers for bacterial infection events and nitrogen fixation.
Newly designed constructs are built using synthetic biology and golden gate cloning. Subsequently, the functionality of the constructs is tested in legume models (e.g. Medicago truncatula and Lotus japonicus), the nodulating non-legume Parasponia, and non-nodulating sister species that have lost the nodulation in course of evolution. To do so, fast and efficient transformation protocols have been established, allowing a ‘design-built-test’ cycle within ~3 months.
In a research thesis you will come familiar with this ‘design-built-test’ cycle and all techniques which are associated with it.
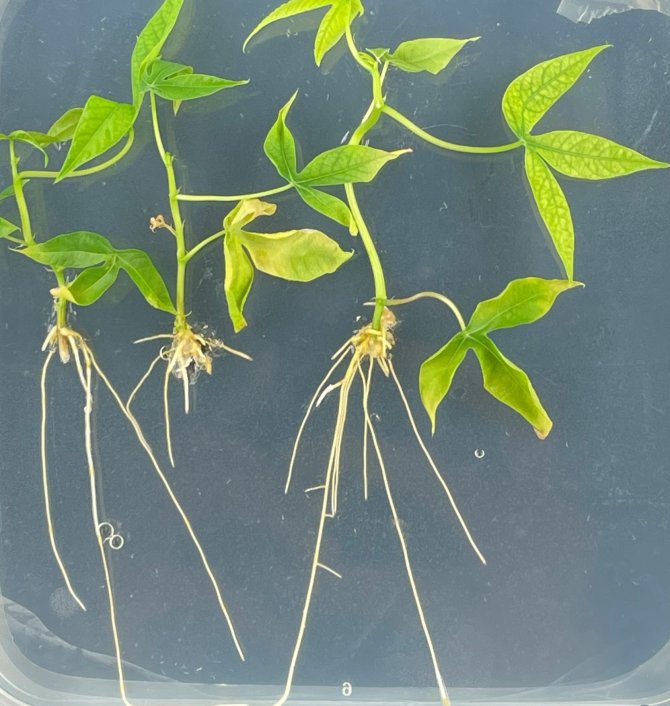
Engineering cassava
Cassava as an important carbon-rich staple food for millions of people in West-Africa and other regions in the world. We selected cassava as an engineering target crop because the species is phylogenetically closely related to the so-called nitrogen-fixation clade. The nitrogen-fixation clade represents all plant species that can form root nodules in symbiosis either with rhizobium bacteria or Frankia bacteria. Current hypothesis on root nodulation in the nitrogen-fixing clade suggests a single evolutionary origin. The cassava ancestor just missed this opportunity!
We established a fast Agrobecterium rhizogenes-mediated root transformation protocol for cassava. This protocol allows bypassing the tedious Agrobacterium tumefaciens transformation procedure that is generally used for cassava. By using the A. rhizogenes transformation protocol, cassava plants carrying transgenic roots can be obtained in 4 weeks, allowing fast testing of engineering constructs.
In a research thesis you will come familiar with cassava biology, root development, transformation and engineering strategy.
