Assembling a minimal, functional actomyosin cortex inside liposomes
The ability of eukaryotic cells to change shape and migrate within 3D environments is governed by a highly dynamic intracellular protein machinery known as the cytoskeleton. In particular, actin filaments form a cross-linked meshwork just underneath the cell membrane. This structure, commonly known as the cell cortex, is closely linked to the membrane and can generate contractile forces, thanks mainly to the presence of myosin motors that traverse along actin filaments. This cytoskeleton-driven force generation regulates the cell shape and helps cells to move in response to external stimuli. How the microscopic cortex organization ultimately brings about cell migration still needs to be fully answered1.
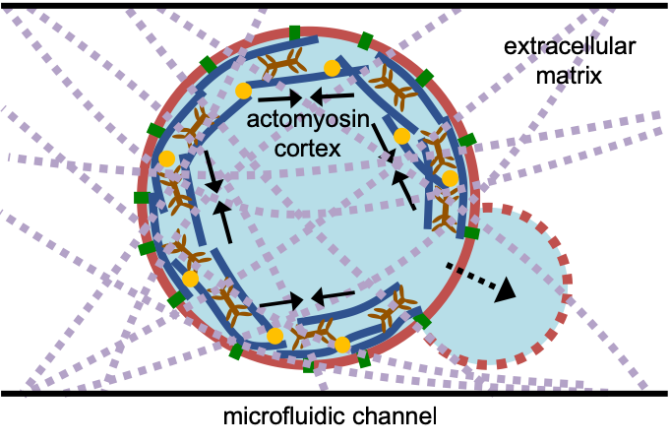
In this exciting project, you will assemble a minimal actomyosin cortex inside liposomes using a creative bottom-up approach. Cell membrane-mimicking liposomes will be produced on a microfluidic chip using Octanol-assisted Liposome Assembly (abbreviated as OLA), a technique that our lab has recently developed2. You will be involved in efficiently encapsulating actin and other essential components inside liposomes and design a system to recruit the cytoskeletal proteins to the membrane, to ultimately form a functional cortex. By stimulating the cortex dynamics through key physiochemical parameters, we will study their effect on liposome shape using fluorescence microscopy.
If you want to get hands-on experience with microfluidics, microscopy, and synthetic biology, just drop us an email (siddharth.deshpande@wur.nl) or pass by our lab/office (Helix, 7056).
References
1. Simon, C. et al. Actin dynamics drive cell-like membrane deformation. Nat. Phys. 15, 602–609 (2019).
2. Deshpande, S. & Dekker, C. On-chip microfluidic production of cell-sized liposomes. Nat. Protoc. 13, 856–874 (2018).