Project
Exploiting the anti-CRISPR potential to enhance the CRISPR genome engineering toolbox
CRISPR/Cas systems are prokaryotic immune systems against bacteriophages and mobile genetic elements. Anti-CRISPR/Cas systems are bacteriophage encoded systems to counterattack CRISPR/Cas immunity. Here, we exploit these systems to develop novel and enhanced genome engineering tools for bacteria.
Background
Bacteriophages (or phages) are viruses that infect bacteria. Phages are the most abundant and diverse biological entities on the planet, exceeding 10-fold over bacterial cells. The evolution of phages has triggered their bacterial hosts to develop an impressive arsenal of defence mechanisms. One of these mechanisms is called CRISPR-Cas (Clustered Regularly Interspaced Short Palindromic Repeats – CRISPR associated proteins).
CRISPR-Cas is a prokaryotic immune system that has been recently repurposed as a next-generation tool for bacterial genome engineering. The popularity of the CRISPR-Cas toolkit emanates from its high efficiency and simplicity. It usually relies on only two elements:
- a Cas9 endonuclease
- a programmable single-guide RNA (sgRNA) molecule
The sgRNA guides Cas9 nuclease to introduce double-stranded DNA breaks to a desired DNA sequence, which is complementary to the exchangeable 5'-end of the sgRNA. In our lab, we recently characterized and exploited novel CRISPR-Cas endonucleases to develop genetic engineering tools for a broad-range of academically and industrially interesting bacteria (mesophiles, thermophiles).
Aim of the project
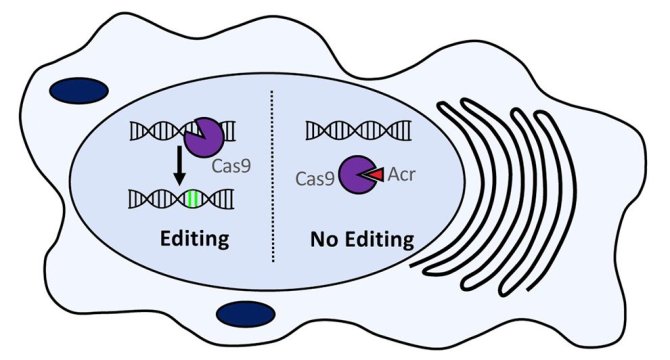
Recently, small proteins naturally inhibiting CRISPR-Cas systems (anti-CRISPR proteins) were identified in phages and mobile genetic elements. These native “off-switches” either block binding of the Cas proteins to their DNA targets or allow binding but prevent cleavage of these targets. Here, we combine anti-CRISPR proteins with novel CRISPR-Cas endonucleases to create highly controllable and efficient tools for bacterial genome editing and silencing.
Techniques
Depending on the project, you will have the possibility to gain expertise on the following techniques: Molecular cloning, protein purification, in vitro transcription, in vitro DNA cleavage & in vivo inhibition assays, flow cytometry analysis, genome editing and silencing in mesophiles/thermophiles.
Contact
Are you an enthusiastic BSc or MSc student that wants to learn more about and work with the cutting-edge CRISPR and anti-CRISPR technologies via a thesis project? Feel free to contact me via raymond.staals@wur.nl.