Project
A viral genome recoded with an unnatural base pair
Next to A, C, T and G, the genetic code can be expanded through the addition of novel molecules, so-called unnatural bases (UBs). This project will generate a UB-containing viral genome, which will be (i) introduced into human host cells to study crucial cellular processes and (ii) used to investigate whether the introduction of UBs can be developed into a biocontainment strategy. The proposed model virus creates a versatile and comprehensive platform to advance the field of synthetic biology.
Background
Xenobiology is the application of unnatural bases (UBs) to supplement the genetic code and expand the space of possibilities for new life functions. This rapidly emerging discipline holds great promise for several fields of science, such as evolutionary studies into the origin of the genetic code. In addition, it can open doors to biotechnological applications such as the creation of novel proteins with functions ranging from materials to therapeutics, and provide a possible biocontainment strategy, as replication of UB-containing genetic material is dependent on the exogenous addition of these UBs (Schmidt, 2010; Budisa et al., 2020).
Thus far, the development of such semi-synthetic organisms (SSO), harboring UBs in their genetic code, remains limited to bacterial systems (Zhang et al., 2017; Hashimoto et al., 2021). Eukaryotic systems are being explored, but complete replication, transcription and translation of UB-containing genes has not been reported (Zhou et al., 2019; Oh et al., 2021). To facilitate the use of UBs in eukaryotic cells, a UB-containing virus can be used to independently study these cellular processes. Viruses are small genetic entities that rely on their host cell for their replication. Their small genome sizes and the availability of extensive genetic toolboxes provide a high degree of flexibility for the introduction of strategically placed UB-containing reporter genes that can be investigated independently.
Project description
This proof-of-principle project uses the well-studied alphavirus Venezuelan equine encephalitis virus (VEEV) to introduce UBs into the eukaryotic cell. It aims to create a VEEV replicon system that encodes a UB-containing reporter gene on a separate ‘minigenome’. This presents an adaptable, versatile and innovative platform to tease apart RNA translation, transcription and replicational processes within eukaryotic cells.
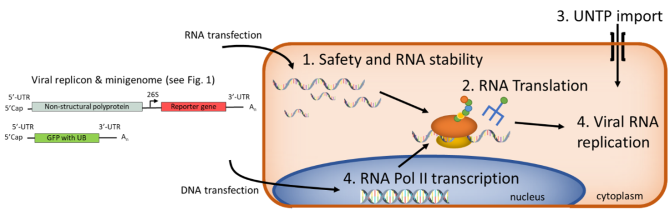
Minigenomes with UBs will be synthesized and transfected with and without the VEEV replicon into human cells to determine stability and retention of the (semi-synthetic) RNA and give a first indication of biosafety (1). This template can be directly translated by the cellular machinery upon addition of appropriate tRNAs to measure protein production (2). Next, we will investigate two strategies to achieve active and prolonged import of unnatural nucleoside triphosphates (UNTPs) via the expression of known nucleoside triphosphate transporters and novel synthetic molecular transporters (3). Finally, we test RNA transcription of the UB-containing minigenome from a DNA template by cellular RNA polymerase II and RNA replication by the VEEV RNA dependent RNA polymerase. The project is embedded in a safe-by-design framework, which aims to account for safety and societal implications at all stages.