
Molecular development of Arbuscular Mycorrhizal symbiosis (Limpens Group)
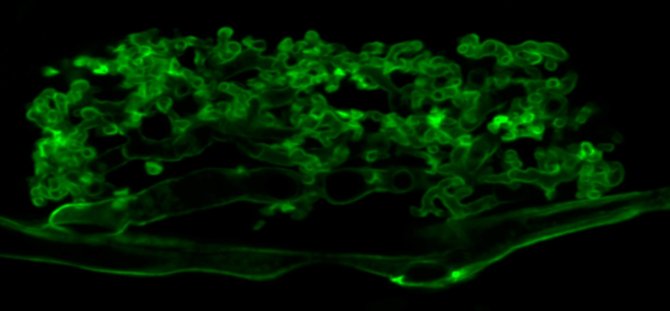
To live in environments where nutrients are limited, plants engage in an endosymbiosis with arbuscular mycorrhizal (AM) fungi. These fungi colonize plant roots and are hosted inside root cortex cells, where highly branched hyphal structures called arbuscules are formed (Figure 1).
There, the fungi deliver scarce minerals, especially phosphate and nitrogen sources, that they take up from the soil to the plant for which they get sugars and lipids in return. The AM symbiosis is very ancient and occurs in the vast majority of all land plants, which makes it one of the agriculturally and ecologically most important endosymbioses in plants.
Understanding how AM fungi are accommodated inside plant roots and how nutrient exchange is controlled is of major importance as it determines the symbiotic efficiency of the interaction. Therefore, we use molecular, genetic- and cell-biological approaches to study the molecular mechanisms by which AM fungi optimize symbiotic nutrient transfer; both from the plant and the fungal side.For more information of specific topics please contact me:
Thesis subjects can be done in the following projects:
How is arbuscule development regulated?
The ability to host AM fungi inside cells requires a reprogramming of root cortex cells, which is controlled by a signalling cascade initiated between fungus and plant. Several key factors controlling arbuscule formation, branching and maintenance (lifetime) have been identified. To unravel the process of arbuscule development we use cell-type specific (transcriptome) analyses to identify novel key regulators and their interactors. A major challenge will be to unravel how these factors work together to control arbuscule development. Currently, we are focussing among others on the role of communication between endodermis and cortex to control arbuscule development and the role of phosphate-sensing transcriptional regulators in the regulation of arbuscule lifetime.
In this research reverse genetic analyses (CRISPR/RNAi/transposon-tagged lines), confocal fluorescence microscopy, protein-protein interactions (co-IP, mass-spec, Y2H) and cell-specific transcriptome analyses (in situ, reporter constructs, qPCR and RNAseq) are used to get insight into the molecular processes that control arbuscule development.
How does genetic variation within fungal isolates contribute to symbiotic efficiency?
One of the key questions in AM biology is to understand at a molecular level what determines how much growth benefit the plant has from interacting with AM fungi. It has been shown that some fungi improve plant growth much more than others. Such variation is not only observed between AMF species or isolates of the same species but even within fungal individuals. However, the molecular basis for such functional diversity is still largely unknown. We try to study how genetic variation between nuclei that populate the common hyphal network (Figure 3) is distributed, their dynamics and ultimately how it affects plant performance. A major challenge for the future will be to find ways to genetically manipulate the fungus.
In this project we combine next-generation sequencing with single cell, spore and single nucleus analyses. Nanotechnology will be pioneered (together with Viola Willemsen) to manipulate the fungus. In addition to wet lab experiments, the project also offers a possibility to focus more on bioinformatics analyses of next-gen sequencing data.
How does the hyphosphere microbiome influence AM symbiosis?
In this novel project, a with collaboration the Chinese Agricultural University (prof. Feng Gu and dr. Lin Zhang), we aim to study how AM fungi work together with bacteria in the soil to facilitate their interaction with the plant; ie. a tripartite interaction between plant-fungus-bacteria. It has become clear that AM fungi can recruit specific bacteria to their extraradical hyphae, which can facilitate the uptake of organic nutrients. We aim to study how bacteria are recruited, the influence of fungal or plant genotypes on which bacteria are recruited and what their role is in the efficiency of the AM interaction with the plant.
In this project, we will use high throughput comparative DNA (amplicon) sequencing for identification of bacteria in combination with meta-transcriptome sequencing. Isolated bacteria will be characterized and their influence on fungus and plant will be studied depending on predictions from bioassays and transcriptome data. Additionally, the role of fungal effector proteins on hyphosphere composition will be studied.